| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-09-30 22:19:15 UTC |
|---|
| Update Date | 2020-05-11 20:46:18 UTC |
|---|
| BMDB ID | BMDB0000070 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
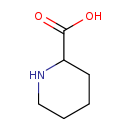
| Common Name | Pipecolic acid |
|---|
| Description | DL-Pipecolic acid, also known as pipecolinate or homoproline, belongs to the class of organic compounds known as alpha amino acids. These are amino acids in which the amino group is attached to the carbon atom immediately adjacent to the carboxylate group (alpha carbon). DL-Pipecolic acid exists as a solid, possibly soluble (in water), and a very strong basic compound (based on its pKa) molecule. DL-Pipecolic acid exists in all living species, ranging from bacteria to humans. DL-Pipecolic acid has been found to be associated with several diseases known as malaria, schizophrenia, rhizomelic chondrodysplasia punctata, and perillyl alcohol administration for cancer treatment; also dl-pipecolic acid has been linked to the inborn metabolic disorders including adrenoleukodystrophy. |
|---|
| Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 2-Piperidinecarboxylic acid | ChEBI | | Homoproline | ChEBI | | Pipecolinic acid | ChEBI | | 2-Piperidinecarboxylate | Generator | | Pipecolinate | Generator | | Pipecolate | Generator | | ()-Piperidine-2-carboxylic acid | HMDB | | (+/-)-2-piperidinecarboxylate | HMDB | | (+/-)-2-piperidinecarboxylic acid | HMDB | | (+/-)-pipecolate | HMDB | | (+/-)-pipecolic acid | HMDB | | (+/-)-pipecolinate | HMDB | | (+/-)-pipecolinic acid | HMDB | | (.+/-.)-2-piperidinecarboxylic acid | HMDB | | (RS)-2-Piperidinecarboxylate | HMDB | | (RS)-2-Piperidinecarboxylic acid | HMDB | | .alpha.-pipecolinic acid | HMDB | | 2-Carboxypiperidine | HMDB | | 2-Pipecolinic acid | HMDB | | 2-Piperidinylcarboxylic acid | HMDB | | a-Pipecolinate | HMDB | | a-Pipecolinic acid | HMDB | | Acide pipecolique | HMDB | | Acide piperidine-carboxylique-2 | HMDB | | alpha-Pipecolinate | HMDB | | alpha-Pipecolinic acid | HMDB | | Dihydrobaikiane | HMDB | | DL-2-Piperidinecarboxylate | HMDB | | DL-2-Piperidinecarboxylic acid | HMDB | | DL-Homoproline | HMDB | | DL-Pipecolate | HMDB, Generator | | DL-Pipecolic acid | HMDB | | DL-Pipecolinate | HMDB | | DL-Pipecolinic acid | HMDB | | hexahydro-2-Picolinate | HMDB | | hexahydro-2-Picolinic acid | HMDB | | Hexahydropicolinate | HMDB | | Hexahydropicolinic acid | HMDB | | Pipecolic acid free base | HMDB | | Piperidine-2-carboxylic acid | HMDB | | Piperolinate | HMDB | | Piperolinic acid | HMDB | | L-Pipecolic acid | MeSH, HMDB | | Pipecolic acid, (R)-isomer | MeSH, HMDB | | Pipecolic acid, ion (1-) | MeSH, HMDB | | Pipecolic acid, monopotassium salt | MeSH, HMDB | | Pipecolic acid hydrochloride, (+-)-isomer | MeSH, HMDB | | Pipecolic acid, (S)-isomer | MeSH, HMDB | | Pipecolic acid, 14C-labeled CPD, (+,-)-isomer | MeSH, HMDB | | Homopipecolic acid | MeSH, HMDB | | Pipecolic acid, (+,-)-isomer | MeSH, HMDB | | Pipecolic acid, ion(1-), (+,-)-isomer | MeSH, HMDB | | Pipecolic acid, ion(1-), (S)-isomer | MeSH, HMDB | | Piperidine-2-carboxylate | Generator, HMDB | | Pipecolic acid | MeSH |
|
|---|
| Chemical Formula | C6H11NO2 |
|---|
| Average Molecular Weight | 129.157 |
|---|
| Monoisotopic Molecular Weight | 129.078978601 |
|---|
| IUPAC Name | piperidine-2-carboxylic acid |
|---|
| Traditional Name | (+,-)-pipecolic acid |
|---|
| CAS Registry Number | 535-75-1 |
|---|
| SMILES | OC(=O)C1CCCCN1 |
|---|
| InChI Identifier | InChI=1S/C6H11NO2/c8-6(9)5-3-1-2-4-7-5/h5,7H,1-4H2,(H,8,9) |
|---|
| InChI Key | HXEACLLIILLPRG-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as alpha amino acids. These are amino acids in which the amino group is attached to the carbon atom immediately adjacent to the carboxylate group (alpha carbon). |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic acids and derivatives |
|---|
| Class | Carboxylic acids and derivatives |
|---|
| Sub Class | Amino acids, peptides, and analogues |
|---|
| Direct Parent | Alpha amino acids |
|---|
| Alternative Parents | |
|---|
| Substituents | - Alpha-amino acid
- Piperidinecarboxylic acid
- Piperidine
- Amino acid
- Azacycle
- Organoheterocyclic compound
- Secondary amine
- Monocarboxylic acid or derivatives
- Secondary aliphatic amine
- Carboxylic acid
- Hydrocarbon derivative
- Organopnictogen compound
- Organic oxygen compound
- Organooxygen compound
- Organonitrogen compound
- Carbonyl group
- Amine
- Organic nitrogen compound
- Organic oxide
- Aliphatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aliphatic heteromonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Status | Detected but not Quantified |
|---|
| Origin | |
|---|
| Biofunction | Not Available |
|---|
| Application | Not Available |
|---|
| Cellular locations | |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Properties | | Property | Value | Reference |
|---|
| Melting Point | 264 °C | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | 314 mg/mL | Not Available | | LogP | -2.31 | TSAI,RS ET AL. (1991) |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | |
|---|
| GC-MS | GC-MS Spectrum - GC-EI-TOF (Non-derivatized) | splash10-0a4i-0910000000-d9cfd443adb5187a3f21 | View in MoNA |
|---|
| GC-MS | GC-MS Spectrum - GC-EI-TOF (Non-derivatized) | splash10-0a4i-0910000000-d9cfd443adb5187a3f21 | View in MoNA |
|---|
| GC-MS | GC-MS Spectrum - GC-EI-TOF (Non-derivatized) | splash10-0a4i-0910000000-d9cfd443adb5187a3f21 | View in MoNA |
|---|
| GC-MS | GC-MS Spectrum - GC-EI-TOF (Non-derivatized) | splash10-0a4i-0910000000-d9cfd443adb5187a3f21 | View in MoNA |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-001i-9200000000-5c4f6059bcc868ca23fd | View in MoNA |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | splash10-05fu-9700000000-8003521353d4d37af49f | View in MoNA |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_2) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_1) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_2) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 10V, Positive (Annotated) | splash10-0060-9900000000-c5b5bf28424d5d5d1bda | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 25V, Positive (Annotated) | splash10-001i-9000000000-92660ad26bff36fba7ef | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 40V, Positive (Annotated) | splash10-0a59-9000000000-b779372fa816467754c1 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF (UPLC Q-Tof Premier, Waters) , Positive | splash10-001i-9300000000-26dee47121401990f1c1 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF (UPLC Q-Tof Premier, Waters) 30V, Positive | splash10-001i-9000000000-929057dfd61546bee24d | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF (UPLC Q-Tof Premier, Waters) , Negative | splash10-004i-0900000000-6595801dc6a322f712f0 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , negative | splash10-004i-0900000000-6595801dc6a322f712f0 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - , negative | splash10-004i-0900000000-99a7a942cee58c76115a | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-001i-9300000000-26dee47121401990f1c1 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-001i-9000000000-929057dfd61546bee24d | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - , positive | splash10-001i-9600000000-9631d0fbc984509f8d1e | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - 30V, Positive | splash10-001i-9000000000-b244255a58ffb3b92be5 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - 35V, Positive | splash10-001i-9000000000-c3041e7b5b1f5118c1ec | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-001i-3900000000-b2ceb4c88f50a6524604 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-001i-9400000000-d3fab76e3fbfbe92c14d | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a5c-9000000000-20464a5c62992da01ab7 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-004i-2900000000-6fac2bf48ffe4d910118 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-003r-8900000000-81b6d69c7a568e36b49f | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-001i-9000000000-6332d1986eb0da0057fd | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-004i-0900000000-b487cd2a72f17977a8cd | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-01t9-1900000000-83b59287dc6d3e726326 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0036-9200000000-3ca813e7348e686ed922 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-001i-9300000000-cb8df30f7267d610fd8d | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-001i-9000000000-72c8691eb15dce6e0abb | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-053r-9000000000-c7623c5058ad6c879ec3 | View in MoNA |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 500 MHz, H2O, experimental) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 2D NMR | [1H, 1H]-TOCSY. Unexported temporarily by An Chi on Oct 15, 2021 until json or nmrML file is generated. 2D NMR Spectrum (experimental) | Not Available | View in JSpectraViewer |
|---|
| 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 600 MHz, H2O, experimental) | Not Available | View in JSpectraViewer |
|---|
|
|---|
| General References | - Melzer N, Wittenburg D, Hartwig S, Jakubowski S, Kesting U, Willmitzer L, Lisec J, Reinsch N, Repsilber D: Investigating associations between milk metabolite profiles and milk traits of Holstein cows. J Dairy Sci. 2013 Mar;96(3):1521-34. doi: 10.3168/jds.2012-5743. [PubMed:23438684 ]
- Mung D, Li L: Development of Chemical Isotope Labeling LC-MS for Milk Metabolomics: Comprehensive and Quantitative Profiling of the Amine/Phenol Submetabolome. Anal Chem. 2017 Apr 18;89(8):4435-4443. doi: 10.1021/acs.analchem.6b03737. Epub 2017 Mar 28. [PubMed:28306241 ]
|
|---|