| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-09-30 23:31:56 UTC |
|---|
| Update Date | 2020-05-21 16:27:47 UTC |
|---|
| BMDB ID | BMDB0007168 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
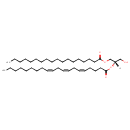
| Common Name | DG(18:0/20:3(5Z,8Z,11Z)/0:0) |
|---|
| Description | DG(18:0/20:3(5Z,8Z,11Z)/0:0) belongs to the family of Diacylglycerols. These are glycerolipids lipids containing a common glycerol backbone to which at least one fatty acyl group is esterified. DG(18:0/20:3(5Z,8Z,11Z)/0:0) is also a substrate of diacylglycerol kinase. It is involved in the phospholipid metabolic pathway. |
|---|
| Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 1-Stearoyl-2-meadoyl-sn-glycerol | HMDB | | DAG(18:0/20:3) | HMDB | | DAG(18:0/20:3N9) | HMDB | | DAG(18:0/20:3W9) | HMDB | | DAG(38:3) | HMDB | | DG(18:0/20:3) | HMDB | | DG(18:0/20:3N9) | HMDB | | DG(18:0/20:3W9) | HMDB | | DG(38:3) | HMDB | | Diacylglycerol | HMDB | | Diacylglycerol(18:0/20:3) | HMDB | | Diacylglycerol(18:0/20:3n9) | HMDB | | Diacylglycerol(18:0/20:3W9) | HMDB | | Diacylglycerol(38:3) | HMDB | | Diglyceride | HMDB | | 1-Octadecanoyl-2-(5Z,8Z,11Z-eicosatrienoyl)-sn-glycerol | HMDB | | DG(18:0/20:3(5Z,8Z,11Z)/0:0) | Lipid Annotator |
|
|---|
| Chemical Formula | C41H74O5 |
|---|
| Average Molecular Weight | 647.0233 |
|---|
| Monoisotopic Molecular Weight | 646.553625478 |
|---|
| IUPAC Name | (2S)-1-hydroxy-3-(octadecanoyloxy)propan-2-yl (5Z,8Z,11Z)-icosa-5,8,11-trienoate |
|---|
| Traditional Name | diacylglycerol |
|---|
| CAS Registry Number | Not Available |
|---|
| SMILES | [H][C@](CO)(COC(=O)CCCCCCCCCCCCCCCCC)OC(=O)CCC\C=C/C\C=C/C\C=C/CCCCCCCC |
|---|
| InChI Identifier | InChI=1S/C41H74O5/c1-3-5-7-9-11-13-15-17-19-20-22-24-26-28-30-32-34-36-41(44)46-39(37-42)38-45-40(43)35-33-31-29-27-25-23-21-18-16-14-12-10-8-6-4-2/h17,19,22,24,28,30,39,42H,3-16,18,20-21,23,25-27,29,31-38H2,1-2H3/b19-17-,24-22-,30-28-/t39-/m0/s1 |
|---|
| InChI Key | UOFQEGAUILMICB-SRRHGZAXSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as 1,2-diacylglycerols. These are diacylglycerols containing a glycerol acylated at positions 1 and 2. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Glycerolipids |
|---|
| Sub Class | Diradylglycerols |
|---|
| Direct Parent | 1,2-diacylglycerols |
|---|
| Alternative Parents | |
|---|
| Substituents | - 1,2-acyl-sn-glycerol
- Fatty acid ester
- Fatty acyl
- Dicarboxylic acid or derivatives
- Carboxylic acid ester
- Carboxylic acid derivative
- Organic oxygen compound
- Organic oxide
- Hydrocarbon derivative
- Primary alcohol
- Organooxygen compound
- Carbonyl group
- Alcohol
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Ontology |
|---|
| Status | Expected but not Quantified |
|---|
| Origin | |
|---|
| Biofunction | Not Available |
|---|
| Application | Not Available |
|---|
| Cellular locations | |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_1) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_1) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS ("DG(18:0/20:3(5Z,8Z,11Z)/0:0),1TMS,#1" TMS) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-03di-0000009000-2259905083239ea1ecd7 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-03fu-0009004000-ee0b2bd938d5c7d97d7b | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-03dl-0009004000-1fa7f2b20eba51aefcb5 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-014i-0000009000-d37410aaac9880d0864d | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-014i-0000009000-d37410aaac9880d0864d | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-03e0-0009000000-f5ccee785f7e2114a45b | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0532-2068009000-81349be214477ab2d1b8 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-001i-2093000000-9585c2109acb806a699f | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0api-3194000000-bf35bf4bd75dd8db5ba6 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-03di-0000009000-63265ec42c7a24551d59 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-03fu-0009004000-15acaa7825d0ce439ad5 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-03dl-0009004000-0842d608eb3662c76705 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-01p2-2088019000-d42cd4aeb098cd9bbfe6 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-000i-3193010000-8f37995cb062a80d0b18 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-06ri-6479000000-949028f1136b51623057 | View in MoNA |
|---|
|
|---|
| Pathways | |
|---|