| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-09-30 23:33:17 UTC |
|---|
| Update Date | 2020-05-21 16:27:52 UTC |
|---|
| BMDB ID | BMDB0007235 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
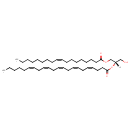
| Common Name | DG(18:1(9Z)/22:5(4Z,7Z,10Z,13Z,16Z)/0:0) |
|---|
| Description | DG(18:1(9Z)/22:5(4Z,7Z,10Z,13Z,16Z)/0:0), also known as dg(18:1(9z)/22:5(4z,7z,10z,13z,16z)/0:0) or DG(18:1/22:5), belongs to the class of organic compounds known as 1,2-dg(18:1(9z)/22:5(4z,7z,10z,13z,16z)/0:0)s. These are dg(18:1(9z)/22:5(4z,7z,10z,13z,16z)/0:0)s containing a glycerol acylated at positions 1 and 2. DG(18:1(9Z)/22:5(4Z,7Z,10Z,13Z,16Z)/0:0) is possibly soluble (in water) and an extremely weak basic (essentially neutral) compound (based on its pKa). DG(18:1(9Z)/22:5(4Z,7Z,10Z,13Z,16Z)/0:0) participates in a number of enzymatic reactions, within cattle. In particular, DG(18:1(9Z)/22:5(4Z,7Z,10Z,13Z,16Z)/0:0) can be biosynthesized from PA(18:1(9Z)/22:5(4Z,7Z,10Z,13Z,16Z)) through the action of the enzyme phosphatidate phosphatase. In addition, DG(18:1(9Z)/22:5(4Z,7Z,10Z,13Z,16Z)/0:0) and tetracosanoyl-CoA can be converted into TG(18:1(9Z)/22:5(4Z,7Z,10Z,13Z,16Z)/24:0); which is mediated by the enzyme dg(18:1(9z)/22:5(4z,7z,10z,13z,16z)/0:0) O-acyltransferase. In cattle, DG(18:1(9Z)/22:5(4Z,7Z,10Z,13Z,16Z)/0:0) is involved in the metabolic pathway called de novo triacylglycerol biosynthesis TG(18:1(9Z)/22:5(4Z,7Z,10Z,13Z,16Z)/24:0) pathway. |
|---|
| Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| DAG(40:6) | HMDB | | DG(18:1/22:5) | HMDB | | 1-Oleoyl-2-osbondoyl-sn-glycerol | HMDB | | Diglyceride | HMDB | | DG(40:6) | HMDB | | Diacylglycerol | HMDB | | Diacylglycerol(18:1/22:5) | HMDB | | DAG(18:1/22:5) | HMDB | | 1-(9Z-Octadecenoyl)-2-(4Z,7Z,10Z,13Z,16Z-docosapentaenoyl)-sn-glycerol | HMDB | | Diacylglycerol(40:6) | HMDB | | DG(18:1(9Z)/22:5(4Z,7Z,10Z,13Z,16Z)/0:0) | Lipid Annotator |
|
|---|
| Chemical Formula | C43H72O5 |
|---|
| Average Molecular Weight | 669.0288 |
|---|
| Monoisotopic Molecular Weight | 668.537975414 |
|---|
| IUPAC Name | (2S)-1-hydroxy-3-[(9Z)-octadec-9-enoyloxy]propan-2-yl (4Z,7Z,10Z,13Z,16Z)-docosa-4,7,10,13,16-pentaenoate |
|---|
| Traditional Name | diacylglycerol |
|---|
| CAS Registry Number | Not Available |
|---|
| SMILES | [H][C@](CO)(COC(=O)CCCCCCC\C=C/CCCCCCCC)OC(=O)CC\C=C/C\C=C/C\C=C/C\C=C/C\C=C/CCCCC |
|---|
| InChI Identifier | InChI=1S/C43H72O5/c1-3-5-7-9-11-13-15-17-19-20-21-22-24-26-28-30-32-34-36-38-43(46)48-41(39-44)40-47-42(45)37-35-33-31-29-27-25-23-18-16-14-12-10-8-6-4-2/h11,13,17-19,21-23,26,28,32,34,41,44H,3-10,12,14-16,20,24-25,27,29-31,33,35-40H2,1-2H3/b13-11-,19-17-,22-21-,23-18-,28-26-,34-32-/t41-/m0/s1 |
|---|
| InChI Key | ZSUNRJOJJXRACA-PYQPTFFQSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as 1,2-diacylglycerols. These are diacylglycerols containing a glycerol acylated at positions 1 and 2. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Glycerolipids |
|---|
| Sub Class | Diradylglycerols |
|---|
| Direct Parent | 1,2-diacylglycerols |
|---|
| Alternative Parents | |
|---|
| Substituents | - 1,2-acyl-sn-glycerol
- Fatty acid ester
- Fatty acyl
- Dicarboxylic acid or derivatives
- Carboxylic acid ester
- Carboxylic acid derivative
- Organic oxygen compound
- Organic oxide
- Hydrocarbon derivative
- Primary alcohol
- Organooxygen compound
- Carbonyl group
- Alcohol
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Ontology |
|---|
| Status | Expected but not Quantified |
|---|
| Origin | |
|---|
| Biofunction | Not Available |
|---|
| Application | Not Available |
|---|
| Cellular locations | |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-01p9-3297032000-496e42c946a8f9790263 | View in MoNA |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_1) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_1) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-040r-0096003000-d5a69e8e6dd6e3fceb72 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-01q9-1093000000-0ac4c978fe9d63d0f690 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-03gi-4095000000-a0c4324bf46f7af00f20 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-014i-0036009000-ad7debaeb83df5b9ed72 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-01z9-4096001000-d237e7b29cb6ed15e566 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-08ir-4195000000-ff28d20a10648795ae76 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-02ti-1049004000-540d63d8fdc983fe1b2f | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-02g9-2098022000-f77211d7184fbbb49354 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0079-1093020000-3c00e47293f29356ed91 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-000i-0000009000-e2329eb25be3f42c3a5d | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-000i-0009004000-871eed3d799a25e9dd54 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-000i-0009004000-9ad5e6a1a2ae2f45727e | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-02t9-2039026000-69b0773eb6062dea75f6 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-03dr-2139000000-04184bf41113c6ef072a | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-03dr-1249000000-e56e08f5b62059d373d1 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0006-0000009000-d8fc180957e9e2b5f8d1 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0006-0000009000-d8fc180957e9e2b5f8d1 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0bt9-0009601000-c530c47d525815613a40 | View in MoNA |
|---|
|
|---|
| Pathways | |
|---|