| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-09-30 23:03:52 UTC |
|---|
| Update Date | 2020-06-04 20:32:38 UTC |
|---|
| BMDB ID | BMDB0003345 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
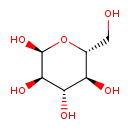
| Common Name | Alpha-D-Glucose |
|---|
| Description | Alpha-D-Glucose, also known as alpha-D-GLC or alphalpha-d-glucose, belongs to the class of organic compounds known as hexoses. These are monosaccharides in which the sugar unit is a is a six-carbon containing moeity. Alpha-D-Glucose exists as a solid, possibly soluble (in water), and an extremely weak basic (essentially neutral) compound (based on its pKa) molecule. Alpha-D-Glucose exists in all living species, ranging from bacteria to humans. Alpha-D-Glucose participates in a number of enzymatic reactions, within cattle. In particular, Alpha-D-Glucose can be converted into glucose 6-phosphate; which is catalyzed by the enzyme glucokinase. In addition, Alpha-D-Glucose and D-fructose can be converted into sucrose through its interaction with the enzyme sucrase-isomaltase, intestinal. In cattle, Alpha-D-glucose is involved in the metabolic pathway called the starch and sucrose metabolism pathway. |
|---|
| Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| alpha-D-GLC | ChEBI | | alpha-Dextrose | ChEBI | | a-D-GLC | Generator | | Α-D-GLC | Generator | | a-Dextrose | Generator | | Α-dextrose | Generator | | a-D-Glucose | Generator | | Α-D-glucose | Generator | | a-D-Glucopyranose | HMDB | | a-Glucose | HMDB | | alpha-D-Glucopyranose | HMDB | | alpha-delta-Glucopyranose | HMDB | | alpha-delta-Glucose | HMDB | | alpha-Glucose | HMDB | | Hexopyranose | HMDB | | Anhydrous dextrose | HMDB | | D Glucose | HMDB | | D-Glucose | HMDB | | Dextrose | HMDB | | Dextrose, anhydrous | HMDB | | Glucose | HMDB | | Glucose monohydrate | HMDB | | Glucose, (DL)-isomer | HMDB | | Glucose, (L)-isomer | HMDB | | Glucose, (alpha-D)-isomer | HMDB | | Glucose, (beta-D)-isomer | HMDB | | L Glucose | HMDB | | L-Glucose | HMDB | | Monohydrate, glucose | HMDB | | 1,3-alpha-D-Glucan | HMDB | | alpha-1,3-Glucan | HMDB | | alpha-D-Glucopyranoside | HMDB |
|
|---|
| Chemical Formula | C6H12O6 |
|---|
| Average Molecular Weight | 180.1559 |
|---|
| Monoisotopic Molecular Weight | 180.063388116 |
|---|
| IUPAC Name | (2S,3R,4S,5S,6R)-6-(hydroxymethyl)oxane-2,3,4,5-tetrol |
|---|
| Traditional Name | α-glucose |
|---|
| CAS Registry Number | 492-62-6 |
|---|
| SMILES | OC[C@H]1O[C@H](O)[C@H](O)[C@@H](O)[C@@H]1O |
|---|
| InChI Identifier | InChI=1S/C6H12O6/c7-1-2-3(8)4(9)5(10)6(11)12-2/h2-11H,1H2/t2-,3-,4+,5-,6+/m1/s1 |
|---|
| InChI Key | WQZGKKKJIJFFOK-DVKNGEFBSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as hexoses. These are monosaccharides in which the sugar unit is a is a six-carbon containing moeity. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic oxygen compounds |
|---|
| Class | Organooxygen compounds |
|---|
| Sub Class | Carbohydrates and carbohydrate conjugates |
|---|
| Direct Parent | Hexoses |
|---|
| Alternative Parents | |
|---|
| Substituents | - Hexose monosaccharide
- Oxane
- Secondary alcohol
- Hemiacetal
- Oxacycle
- Organoheterocyclic compound
- Polyol
- Hydrocarbon derivative
- Primary alcohol
- Alcohol
- Aliphatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aliphatic heteromonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Status | Detected and Quantified |
|---|
| Origin | |
|---|
| Biofunction | Not Available |
|---|
| Application | Not Available |
|---|
| Cellular locations | |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Properties | | Property | Value | Reference |
|---|
| Melting Point | 146 °C | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | 500 mg/mL at 20 °C | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0np0-9700000000-e8d638dc817e46b97d7b | View in MoNA |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (5 TMS) - 70eV, Positive | splash10-004i-6122690000-eaf6f7adf34ccd0c667b | View in MoNA |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - 10V, Negative | splash10-0a4i-9100000000-57ee067515dbaeec1e27 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - 35V, Negative | splash10-0ab9-9100000000-311a61760b7c0dc3a7e1 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - 35V, Negative | splash10-0ab9-9100000000-f2beec08615d57ac1a02 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - 35V, Negative | splash10-0ab9-9100000000-eb694004566014e0b267 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - 20V, Negative | splash10-0a4i-9000000000-906b2fd231873b000e08 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - 40V, Negative | splash10-0a4i-9000000000-3cd19dc7a445df26b86d | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 25V, Positive (Annotated) | splash10-000i-9100000000-53278ff57ca00e1b5d1a | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 40V, Positive (Annotated) | splash10-0079-9000000000-c4182c64988208ac78b0 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-004i-2900000000-a4ec4f0b1e29e360a952 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-01t9-6900000000-7b3ea9c64ecc8d4ac867 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-052f-9100000000-ec2bf4918640a0a36398 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-004i-2900000000-2448926b508622464fe7 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-056r-8900000000-4c073cb93b78120113e6 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0a4l-9000000000-ece70093ab5d3c331ac4 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-01q9-0900000000-b0bc47623e7b2ca31c02 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-03ea-3900000000-648e1637af29cf2a3518 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0007-9200000000-9e6f46a1cbf52d6e347a | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-01qa-0900000000-04ceb34d441ff6a75763 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-03dl-9400000000-a8f1ceab155611f949c1 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-01ow-9000000000-358f68fc2b7a72c27546 | View in MoNA |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 600 MHz, H2O, experimental) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, D2O, experimental) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 600 MHz, H2O, experimental) | Not Available | View in JSpectraViewer |
|---|
|
|---|