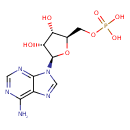
Showing metabocard for Adenosine monophosphate (BMDB0000045)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Creation Date | 2016-09-30 22:18:55 UTC | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 2020-06-04 20:04:56 UTC | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BMDB ID | BMDB0000045 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Secondary Accession Numbers |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Common Name | Adenosine monophosphate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | Adenosine monophosphate, also known as adenylic acid or AMP, belongs to the class of organic compounds known as purine ribonucleoside monophosphates. These are nucleotides consisting of a purine base linked to a ribose to which one monophosphate group is attached. Adenosine monophosphate is a drug which is used for nutritional supplementation, also for treating dietary shortage or imbalance. Adenosine monophosphate exists as a solid, possibly soluble (in water), and a strong basic compound (based on its pKa) molecule. Adenosine monophosphate exists in all living species, ranging from bacteria to humans. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula | C10H14N5O7P | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight | 347.2212 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight | 347.063084339 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name | {[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-3,4-dihydroxyoxolan-2-yl]methoxy}phosphonic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional Name | adenylate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS Registry Number | 61-19-8 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES | NC1=C2N=CN([C@@H]3O[C@H](COP(O)(O)=O)[C@@H](O)[C@H]3O)C2=NC=N1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Identifier | InChI=1S/C10H14N5O7P/c11-8-5-9(13-2-12-8)15(3-14-5)10-7(17)6(16)4(22-10)1-21-23(18,19)20/h2-4,6-7,10,16-17H,1H2,(H2,11,12,13)(H2,18,19,20)/t4-,6-,7-,10-/m1/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key | UDMBCSSLTHHNCD-KQYNXXCUSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | belongs to the class of organic compounds known as purine ribonucleoside monophosphates. These are nucleotides consisting of a purine base linked to a ribose to which one monophosphate group is attached. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Nucleosides, nucleotides, and analogues | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Purine nucleotides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Purine ribonucleotides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Purine ribonucleoside monophosphates | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic heteropolycyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Ontology | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Status | Detected and Quantified | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Origin |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biofunction | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Application | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular locations |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biospecimen Locations |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Normal Concentrations | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Abnormal Concentrations | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HMDB ID | HMDB0000045 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| DrugBank ID | DB00131 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Phenol Explorer Compound ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| FooDB ID | FDB030677 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KNApSAcK ID | C00019347 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemspider ID | 5858 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG Compound ID | C00020 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BioCyc ID | AMP | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BiGG ID | 33534 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikipedia Link | Adenylic_acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| METLIN ID | 5111 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PubChem Compound | 6083 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PDB ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChEBI ID | 16027 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| General References |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Only showing the first 50 proteins. There are 67 proteins in total.
Enzymes
- General function:
- Translation, ribosomal structure and biogenesis
- Specific function:
- Catalyzes the attachment of serine to tRNA(Ser). Is also probably able to aminoacylate tRNA(Sec) with serine, to form the misacylated tRNA L-seryl-tRNA(Sec), which will be further converted into selenocysteinyl-tRNA(Sec).
- Gene Name:
- SARS2
- Uniprot ID:
- Q9N0F3
- Molecular weight:
- 58296.0
Reactions
| L-Serine + Adenosine triphosphate + L-Seryl-tRNA → Adenosine monophosphate + L-Seryl-tRNA(Ser) + Pyrophosphate | details |
- General function:
- Translation, ribosomal structure and biogenesis
- Specific function:
- Catalyzes the attachment of serine to tRNA(Ser) in a two-step reaction: serine is first activated by ATP to form Ser-AMP and then transferred to the acceptor end of tRNA(Ser). Is probably also able to aminoacylate tRNA(Sec) with serine, to form the misacylated tRNA L-seryl-tRNA(Sec), which will be further converted into selenocysteinyl-tRNA(Sec). In the nucleus, binds to the VEGFA core promoter and prevents MYC binding and transcriptional activation by MYC. Recruits SIRT2 to the VEGFA promoter, promoting deacetylation of histone H4 at 'Lys-16' (H4K16). Thereby, inhibits the production of VEGFA and sprouting angiogenesis mediated by VEGFA.
- Gene Name:
- SARS1
- Uniprot ID:
- Q9GMB8
- Molecular weight:
- 58605.0
Reactions
| L-Serine + Adenosine triphosphate + L-Seryl-tRNA → Adenosine monophosphate + L-Seryl-tRNA(Ser) + Pyrophosphate | details |
- General function:
- Nucleotide transport and metabolism
- Specific function:
- Catalyzes a salvage reaction resulting in the formation of AMP, that is energically less costly than de novo synthesis.
- Gene Name:
- APRT
- Uniprot ID:
- Q56JW4
- Molecular weight:
- 19537.0
Reactions
| Adenosine monophosphate + Pyrophosphate → Adenine + Phosphoribosyl pyrophosphate | details |
- General function:
- Carbohydrate transport and metabolism
- Specific function:
- Converts adenosine 3'-phosphate 5'-phosphosulfate (PAPS) to adenosine 5'-phosphosulfate (APS) and 3'(2')-phosphoadenosine 5'- phosphate (PAP) to AMP. Has 1000-fold lower activity towards inositol 1,4-bisphosphate (Ins(1,4)P2) and inositol 1,3,4-trisphosphate (Ins(1,3,4)P3), but does not hydrolyze Ins(1)P, Ins(3,4)P2, Ins(1,3,4,5)P4 or InsP6 (By similarity).
- Gene Name:
- BPNT1
- Uniprot ID:
- Q3ZCK3
- Molecular weight:
- 33328.0
- General function:
- Nucleotide transport and metabolism
- Specific function:
- One of the enzymes of the urea cycle, the metabolic pathway transforming neurotoxic amonia produced by protein catabolism into inocuous urea in the liver of ureotelic animals. Catalyzes the formation of arginosuccinate from aspartate, citrulline and ATP and together with ASL it is responsible for the biosynthesis of arginine in most body tissues.
- Gene Name:
- ASS1
- Uniprot ID:
- P14568
- Molecular weight:
- 46417.0
Reactions
| Adenosine triphosphate + Citrulline + L-Aspartic acid → Adenosine monophosphate + Pyrophosphate + Argininosuccinic acid | details |
- General function:
- Lipid transport and metabolism
- Specific function:
- Catalyzes the synthesis of acetyl-CoA from short-chain fatty acids (By similarity). Propionate is the preferred substrate but can also utilize acetate and butyrate with a much lower affinity.
- Gene Name:
- ACSS3
- Uniprot ID:
- A7MB45
- Molecular weight:
- 74805.0
Reactions
| Propinol adenylate + Adenosine triphosphate + Pantetheine → Propionyl-CoA + Adenosine monophosphate + Pyrophosphate | details |
| Propinol adenylate → Propionic acid + Adenosine monophosphate | details |
| Caproic acid + Adenosine triphosphate + Coenzyme A → Hexanoyl-CoA + Adenosine monophosphate + Pyrophosphate | details |
- General function:
- Translation, ribosomal structure and biogenesis
- Specific function:
- Catalyzes the attachment of threonine to tRNA(Thr) in a two-step reaction: threonine is first activated by ATP to form Thr-AMP and then transferred to the acceptor end of tRNA(Thr). Also edits incorrectly charged tRNA(Thr) via its editing domain, at the post-transfer stage.
- Gene Name:
- TARS1
- Uniprot ID:
- Q3ZBV8
- Molecular weight:
- 83492.0
Reactions
| Adenosine triphosphate + L-Threonine + tRNA(Thr) → Adenosine monophosphate + Pyrophosphate + L-Threonyl-tRNA(Thr) | details |
- General function:
- Involved in magnesium ion binding
- Specific function:
- 3'-exoribonuclease that has a preference for poly(A) tails of mRNAs, thereby efficiently degrading poly(A) tails. Exonucleolytic degradation of the poly(A) tail is often the first step in the decay of eukaryotic mRNAs and is also used to silence certain maternal mRNAs translationally during oocyte maturation and early embryonic development. Involved in nonsense-mediated mRNA decay, a critical process of selective degradation of mRNAs that contain premature stop codons. Also involved in degradation of inherently unstable mRNAs that contain AU-rich elements (AREs) in their 3'-UTR, possibly via its interaction with KHSRP. Probably mediates the removal of poly(A) tails of AREs mRNAs, which constitutes the first step of destabilization (By similarity). Interacts with both the 3'-end poly(A) tail and the 5'-end cap structure during degradation, the interaction with the cap structure being required for an efficient degradation of poly(A) tails (By similarity) (PubMed:10698948, PubMed:9736620). Also able to recognize poly(A) tails of microRNAs such as MIR21 and H/ACA box snoRNAs (small nucleolar RNAs) leading to microRNAs degradation or snoRNA increased stability (By similarity).
- Gene Name:
- PARN
- Uniprot ID:
- P69341
- Molecular weight:
- 73182.0
- General function:
- Nucleotide transport and metabolism
- Specific function:
- Catalyzes the synthesis of phosphoribosylpyrophosphate (PRPP) that is essential for nucleotide synthesis.
- Gene Name:
- PRPS1
- Uniprot ID:
- Q2HJ58
- Molecular weight:
- 34834.0
Reactions
| D-Ribose 5-phosphate + Adenosine triphosphate → Phosphoribosyl pyrophosphate + Adenosine monophosphate | details |
- General function:
- Lipid transport and metabolism
- Specific function:
- Catalyzes the activation of fatty acids by CoA to produce an acyl-CoA, the first step in fatty acid metabolism (PubMed:11382754, PubMed:10561077). Capable of activating medium-chain fatty acids (e.g. butyric (C4) to decanoic (C10) acids), and certain carboxylate-containing xenobiotics, e.g. benzoate (PubMed:10561077, PubMed:11382754). Also catalyzes the activation of lipoate to lipoyl-nucleoside monophosphate (PubMed:11382754). Activates lipoate with GTP at a 1000-fold higher rate than with ATP and activates both (R)- and (S)-lipoate to the respective lipoyl-GMP, with a preference for (R)-lipoate (PubMed:11382754).
- Gene Name:
- ACSM1
- Uniprot ID:
- Q9BEA2
- Molecular weight:
- 64923.0
Reactions
| Dodecanoic acid + Adenosine triphosphate + Coenzyme A → Lauroyl-CoA + Adenosine monophosphate + Pyrophosphate | details |
- General function:
- Replication, recombination and repair
- Specific function:
- Hydrolyzes NAD(P)H to NMNH and AMP (2',5'-ADP), and diadenosine diphosphate to AMP. Has also activity towards NAD(P)(+), ADP-ribose and diadenosine triphosphate. May act to regulate the concentration of peroxisomal nicotinamide nucleotide cofactors required for oxidative metabolism in this organelle.
- Gene Name:
- NUDT12
- Uniprot ID:
- Q29RH3
- Molecular weight:
- 50119.0
Reactions
| NAD + Water → Nicotinamide ribotide + Adenosine monophosphate | details |
| Nicotinic acid adenine dinucleotide + Water → Nicotinic acid mononucleotide + Adenosine monophosphate | details |
- General function:
- Translation, ribosomal structure and biogenesis
- Specific function:
- Catalyzes the specific attachment of an amino acid to its cognate tRNA in a 2 step reaction: the amino acid (AA) is first activated by ATP to form AA-AMP and then transferred to the acceptor end of the tRNA.
- Gene Name:
- DARS1
- Uniprot ID:
- Q3SYZ4
- Molecular weight:
- 57036.0
Reactions
| Adenosine triphosphate + L-Aspartic acid + tRNA(Asp) → Adenosine monophosphate + Pyrophosphate + L-Aspartyl-tRNA(Asp) | details |
- General function:
- Nucleotide transport and metabolism
- Specific function:
- Involved in maintaining the homeostasis of cellular nucleotides by catalyzing the interconversion of nucleoside phosphates. Has GTP:AMP phosphotransferase and ITP:AMP phosphotransferase activities.
- Gene Name:
- AK3
- Uniprot ID:
- P08760
- Molecular weight:
- 25671.0
- General function:
- Amino acid transport and metabolism
- Specific function:
- Synthesizes selenophosphate from selenide and ATP.
- Gene Name:
- SEPHS1
- Uniprot ID:
- Q0VC82
- Molecular weight:
- 42881.0
Reactions
| Adenosine triphosphate + Hydrogen selenide + Water → Adenosine monophosphate + Phosphoroselenoic acid + Hydrogen phosphate | details |
- General function:
- Coenzyme transport and metabolism
- Specific function:
- Catalyzes the ATP-dependent amidation of deamido-NAD to form NAD. Uses L-glutamine as a nitrogen source.
- Gene Name:
- NADSYN1
- Uniprot ID:
- Q3ZBF0
- Molecular weight:
- 79400.0
Reactions
| Nicotinic acid adenine dinucleotide + Adenosine triphosphate + Water + L-Glutamine → NAD + Adenosine monophosphate + Pyrophosphate + L-Glutamic acid | details |
- General function:
- Translation, ribosomal structure and biogenesis
- Specific function:
- Glutamine--tRNA ligase. Plays a critical role in brain development.
- Gene Name:
- QARS1
- Uniprot ID:
- Q3MHH4
- Molecular weight:
- 87643.0
Reactions
| L-Glutamine + Adenosine triphosphate + tRNA(Gln) → Adenosine monophosphate + Pyrophosphate + L-Glutaminyl-tRNA(Gln) | details |
- General function:
- Translation, ribosomal structure and biogenesis
- Specific function:
- T1-TrpRS has aminoacylation activity while T2-TrpRS lacks it. T1-TrpRS and T2-TrpRS possess angiostatic activity. T2-TrpRS inhibits fluid shear stress-activated responses of endothelial cells. Regulates ERK, Akt, and eNOS activation pathways that are associated with angiogenesis, cytoskeletal reorganization and shear stress-responsive gene expression (By similarity).
- Gene Name:
- WARS1
- Uniprot ID:
- P17248
- Molecular weight:
- 53812.0
Reactions
| Adenosine triphosphate + L-Tryptophan + tRNA(Trp) → Adenosine monophosphate + Pyrophosphate + L-Tryptophyl-tRNA(Trp) | details |
- General function:
- Translation, ribosomal structure and biogenesis
- Specific function:
- Mitochondrial aminoacyl-tRNA synthetase that activate and transfer the amino acids to their corresponding tRNAs during the translation of mitochondrial genes and protein synthesis.
- Gene Name:
- WARS2
- Uniprot ID:
- Q3T099
- Molecular weight:
- 40205.0
Reactions
| Adenosine triphosphate + L-Tryptophan + tRNA(Trp) → Adenosine monophosphate + Pyrophosphate + L-Tryptophyl-tRNA(Trp) | details |
- General function:
- Nucleotide transport and metabolism
- Specific function:
- Catalyzes two non-sequential steps in de novo AMP synthesis: converts (S)-2-(5-amino-1-(5-phospho-D-ribosyl)imidazole-4-carboxamido)succinate (SAICAR) to fumarate plus 5-amino-1-(5-phospho-D-ribosyl)imidazole-4-carboxamide, and thereby also contributes to de novo IMP synthesis, and converts succinyladenosine monophosphate (SAMP) to AMP and fumarate.
- Gene Name:
- ADSL
- Uniprot ID:
- A3KN12
- Molecular weight:
- 55484.0
Reactions
| Adenylsuccinic acid → Fumaric acid + Adenosine monophosphate | details |
- General function:
- Coenzyme transport and metabolism
- Specific function:
- Catalyzes the phosphorylation of thiamine to thiamine pyrophosphate. Can also catalyze the phosphorylation of pyrithiamine to pyrithiamine pyrophosphate (By similarity).
- Gene Name:
- TPK1
- Uniprot ID:
- Q5E9T4
- Molecular weight:
- 27028.0
Reactions
| Adenosine triphosphate + Thiamine → Adenosine monophosphate + Thiamine pyrophosphate | details |
- General function:
- Lipid transport and metabolism
- Specific function:
- Catalyzes the conversion of fatty acids such as long-chain and very long-chain fatty acids to their active form acyl-CoAs for both synthesis of cellular lipids, and degradation via beta-oxidation. Can activate diverse saturated, monosaturated and polyunsaturated fatty acids.
- Gene Name:
- ACSBG1
- Uniprot ID:
- Q2KHW5
- Molecular weight:
- 80585.0
Reactions
| Palmitic acid + Adenosine triphosphate + Coenzyme A → Palmityl-CoA + Adenosine monophosphate + Pyrophosphate | details |
| Stearic acid + Coenzyme A + Adenosine triphosphate → Stearoyl-CoA + Pyrophosphate + Adenosine monophosphate | details |
- General function:
- Translation, ribosomal structure and biogenesis
- Specific function:
- Catalyzes the ATP-dependent ligation of histidine to the 3'-end of its cognate tRNA, via the formation of an aminoacyl-adenylate intermediate (His-AMP). Plays a role in axon guidance.
- Gene Name:
- HARS1
- Uniprot ID:
- Q2KI84
- Molecular weight:
- 57285.0
Reactions
| Adenosine triphosphate + L-Histidine + tRNA(His) → Adenosine monophosphate + Pyrophosphate + L-Histidyl-tRNA(His) | details |
- General function:
- Translation, ribosomal structure and biogenesis
- Specific function:
- Mitochondrial aminoacyl-tRNA synthetase that catalyzes the ATP-dependent ligation of histidine to the 3'-end of its cognate tRNA, via the formation of an aminoacyl-adenylate intermediate (His-AMP).
- Gene Name:
- HARS2
- Uniprot ID:
- A5D7V9
- Molecular weight:
- 56914.0
Reactions
| Adenosine triphosphate + L-Histidine + tRNA(His) → Adenosine monophosphate + Pyrophosphate + L-Histidyl-tRNA(His) | details |
- General function:
- Nucleotide transport and metabolism
- Specific function:
- Catalyzes the reversible transfer of the terminal phosphate group between ATP and AMP. Also displays broad nucleoside diphosphate kinase activity. Plays an important role in cellular energy homeostasis and in adenine nucleotide metabolism.
- Gene Name:
- AK1
- Uniprot ID:
- P00570
- Molecular weight:
- 21664.0
Reactions
| Adenosine triphosphate + Adenosine monophosphate →2 ADP | details |
- General function:
- Translation, ribosomal structure and biogenesis
- Specific function:
- Catalyzes the specific attachment of an amino acid to its cognate tRNA in a 2 step reaction: the amino acid (AA) is first activated by ATP to form AA-AMP and then transferred to the acceptor end of the tRNA. Plays a role in the synthesis of ribosomal RNA in the nucleolus.
- Gene Name:
- MARS1
- Uniprot ID:
- Q2T9L8
- Molecular weight:
- 100634.0
Reactions
| Selenomethionine + Adenosine triphosphate + tRNA(Met) → Selenomethionyl-tRNA(Met) + Adenosine monophosphate + Pyrophosphate | details |
| L-Methionine + Adenosine triphosphate + tRNA(Met) → L-Methionyl-tRNA(Met) + Adenosine monophosphate + Pyrophosphate | details |
- General function:
- Involved in G-protein coupled receptor protein signalin
- Specific function:
- Receptor for prostaglandin D2 (PGD2). The activity of this receptor is mainly mediated by G(s) proteins that stimulate adenylate cyclase, resulting in an elevation of intracellular cAMP. A mobilization of calcium is also observed, but without formation of inositol 1,4,5-trisphosphate (By similarity).
- Gene Name:
- PTGDR
- Uniprot ID:
- A5D7K8
- Molecular weight:
- 40372.0
- General function:
- Involved in G-protein coupled receptor activity
- Specific function:
- Receptor for prostacyclin (prostaglandin I2 or PGI2). The activity of this receptor is mediated by G(s) proteins which activate adenylate cyclase.
- Gene Name:
- PTGIR
- Uniprot ID:
- P79393
- Molecular weight:
- 41247.0
- General function:
- Involved in prostaglandin E receptor activity
- Specific function:
- Receptor for prostaglandin E2 (PGE2). The activity of this receptor is mediated by G(s) proteins that stimulate adenylate cyclase. Has a relaxing effect on smooth muscle. May play an important role in regulating renal hemodynamics, intestinal epithelial transport, adrenal aldosterone secretion, and uterine function (By similarity).
- Gene Name:
- PTGER4
- Uniprot ID:
- Q8MJ08
- Molecular weight:
- 53298.0
- General function:
- Not Available
- Specific function:
- Receptor for prostaglandin E2 (PGE2) (PubMed:8396726). The various isoforms have identical ligand binding properties but interact with different second messenger systems: isoform EP3A couples to G(i)/G(o) proteins; isoform EP3B and isoform EP3C couple to G(s), and isoform EP3D couples to G(i), G(s) and G(p) (PubMed:8396726). Required for normal development of fever in response to pyrinogens, including IL1B, prostaglandin E2 and bacterial lipopolysaccharide (LPS). Required for normal potentiation of platelet aggregation by prostaglandin E2, and thus plays a role in the regulation of blood coagulation. Required for increased HCO3(-) secretion in the duodenum in response to mucosal acidification, and thereby contributes to the protection of the mucosa against acid-induced ulceration. Not required for normal kidney function, normal urine volume and osmolality (By similarity).
- Gene Name:
- PTGER3
- Uniprot ID:
- P34979
- Molecular weight:
- 46362.0
- General function:
- Involved in vasopressin receptor activity
- Specific function:
- Receptor for arginine vasopressin (PubMed:7698346, PubMed:8257689). The activity of this receptor is mediated by G proteins which activate adenylate cyclase. Involved in renal water reabsorption (By similarity).
- Gene Name:
- AVPR2
- Uniprot ID:
- P48044
- Molecular weight:
- 40236.0
- General function:
- Translation, ribosomal structure and biogenesis
- Specific function:
- Catalyzes the attachment of alanine to tRNA(Ala) in a two-step reaction: alanine is first activated by ATP to form Ala-AMP and then transferred to the acceptor end of tRNA(Ala). Also edits incorrectly charged tRNA(Ala) via its editing domain.
- Gene Name:
- AARS1
- Uniprot ID:
- A6QLT9
- Molecular weight:
- 106655.0
Reactions
| Adenosine triphosphate + L-Alanine + tRNA(Ala) → Adenosine monophosphate + Pyrophosphate + L-Alanyl-tRNA(Ala) | details |
- General function:
- Involved in melanocyte stimulating hormone receptor act
- Specific function:
- Receptor for MSH (alpha, beta) and ACTH. Does not seem to be active with gamma-MSH. The activity of this receptor is mediated by G proteins which activate adenylate cyclase. Could be involved in spermatogenesis.
- Gene Name:
- MC1R
- Uniprot ID:
- P47798
- Molecular weight:
- 34916.0
- General function:
- Involved in vasoactive intestinal polypeptide receptor
- Specific function:
- This is a receptor for PACAP-27 and PACAP-38. The activity of this receptor is mediated by G proteins which activate adenylyl cyclase. May regulate the release of adrenocorticotropin, luteinizing hormone, growth hormone, prolactin, epinephrine, and catecholamine. May play a role in spermatogenesis and sperm motility. Causes smooth muscle relaxation and secretion in the gastrointestinal tract (By similarity).
- Gene Name:
- ADCYAP1R1
- Uniprot ID:
- Q29627
- Molecular weight:
- 58785.0
- General function:
- Involved in alpha2-adrenergic receptor activity
- Specific function:
- Alpha-2 adrenergic receptors mediate the catecholamine-induced inhibition of adenylate cyclase through the action of G proteins. Component of the ATAC complex, a complex with histone acetyltransferase activity on histones H3 and H4 (By similarity).
- Gene Name:
- ADRA2A
- Uniprot ID:
- Q28838
- Molecular weight:
- 49252.0
- General function:
- Not Available
- Specific function:
- The muscarinic acetylcholine receptor mediates various cellular responses, including inhibition of adenylate cyclase, breakdown of phosphoinositides and modulation of potassium channels through the action of G proteins. Primary transducing effect is adenylate cyclase inhibition. Signaling promotes phospholipase C activity, leading to the release of inositol trisphosphate (IP3); this then triggers calcium ion release into the cytosol (By similarity).
- Gene Name:
- CHRM2
- Uniprot ID:
- P41985
- Molecular weight:
- 51613.0
- General function:
- Involved in G-protein coupled receptor protein signalin
- Specific function:
- Alpha-2 adrenergic receptors mediate the catecholamine-induced inhibition of adenylate cyclase through the action of G proteins.
- Gene Name:
- ADRA2B
- Uniprot ID:
- O77700
- Molecular weight:
- 42840.0
- General function:
- Involved in muscarinic acetylcholine receptor activity
- Specific function:
- The muscarinic acetylcholine receptor mediates various cellular responses, including inhibition of adenylate cyclase, breakdown of phosphoinositides and modulation of potassium channels through the action of G proteins. Primary transducing effect is Pi turnover.
- Gene Name:
- CHRM3
- Uniprot ID:
- P41984
- Molecular weight:
- 66103.0
- General function:
- Involved in lutropin-choriogonadotropic hormone recepto
- Specific function:
- Receptor for lutropin-choriogonadotropic hormone. The activity of this receptor is mediated by G proteins which activate adenylate cyclase.
- Gene Name:
- LHCGR
- Uniprot ID:
- Q28005
- Molecular weight:
- 78456.0
- General function:
- Involved in protein-hormone receptor activity
- Specific function:
- Receptor for the thyroid-stimulating hormone (TSH) or thyrotropin. Also acts as a receptor for the heterodimeric glycoprotein hormone (GPHA2:GPHB5) or thyrostimulin. The activity of this receptor is mediated by G proteins which activate adenylate cyclase. Plays a central role in controlling thyroid cell metabolism.
- Gene Name:
- TSHR
- Uniprot ID:
- Q27987
- Molecular weight:
- 86431.0
- General function:
- Involved in adrenocorticotropin receptor activity
- Specific function:
- Receptor for corticotropin (ACTH). This receptor is mediated by G proteins which activate adenylate cyclase (cAMP).
- Gene Name:
- MC2R
- Uniprot ID:
- P34974
- Molecular weight:
- 33293.0
- General function:
- Involved in follicle-stimulating hormone receptor activity
- Specific function:
- G protein-coupled receptor for follitropin, the follicle-stimulating hormone. Through cAMP production activates the downstream PI3K-AKT and ERK1/ERK2 signaling pathways.
- Gene Name:
- FSHR
- Uniprot ID:
- P35376
- Molecular weight:
- 78085.0
- General function:
- Not Available
- Specific function:
- The muscarinic acetylcholine receptor mediates various cellular responses, including inhibition of adenylate cyclase, breakdown of phosphoinositides and modulation of potassium channels through the action of G proteins. Primary transducing effect is inhibition of adenylate cyclase. May couple to multiple functional responses in cell lines.
- Gene Name:
- CHRM4
- Uniprot ID:
- P41986
- Molecular weight:
- 13221.0
- General function:
- Not Available
- Specific function:
- Not Available
- Gene Name:
- Not Available
- Uniprot ID:
- Q9TRE3
- Molecular weight:
- 3427.0
- General function:
- Involved in melanocortin receptor activity
- Specific function:
- Receptor for MSH (alpha, beta and gamma) and ACTH. The activity of this receptor is mediated by G proteins which activate adenylate cyclase. This receptor is a possible mediator of the immunomodulation properties of melanocortins (By similarity).
- Gene Name:
- MC5R
- Uniprot ID:
- P56451
- Molecular weight:
- 36526.0
Only showing the first 50 proteins. There are 67 proteins in total.